 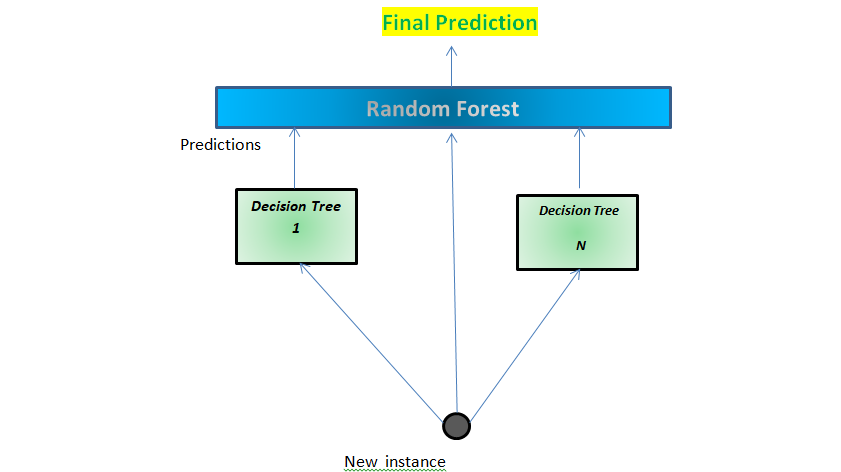
| Generate a bootstrap sample of the original data 5. Select number of trees to build ( ntrees ) 3. The basic algorithm for a regression random forest can be generalized to the following:ġ. When, the randomization amounts to using only step 1 and is the same as bagging. For regression trees, typical default values are but this should be considered a tuning parameter. Split-variable randomization: each time a split is to be performed, the search for the split variable is limited to a random subset of m of the p variables.Bootstrap: similar to bagging, each tree is grown to a bootstrap resampled data set, which makes them different and somewhat decorrelates them.This can be achieved by injecting more randomness into the tree-growing process. In order to reduce variance further, we need to minimize the amount of correlation between the trees. This characteristic is known as tree correlation and prevents bagging from optimally reducing variance of the predictive values. Six decision trees based on different bootstrap samples. Although there are 15 predictor variables to split on, all six trees have both lstat and rm variables driving the first few splits. Rather, trees from different bootstrap samples typically have similar structure to each other (especially at the top of the tree) due to underlying relationships.įor example, if we create six decision trees with different bootstrapped samples of the Boston housing data, we see that the top of the trees all have a very similar structure. However, the trees in bagging are not completely independent of each other since all the original predictors are considered at every split of every tree. Bagging trees introduces a random component in to the tree building process that reduces the variance of a single tree’s prediction and improves predictive performance. Random forests are built on the same fundamental principles as decision trees and bagging (check out this tutorial if you need a refresher on these techniques). 7 ) ames_train <- training ( ames_split ) ames_test <- testing ( ames_split ) The idea # Use set.seed for reproducibility set.seed ( 123 ) ames_split <- initial_split ( AmesHousing :: make_ames (), prop =. # Create training (70%) and test (30%) sets for the AmesHousing::make_ames() data. Some of these packages play a supporting role however, we demonstrate how to implement random forests with several different packages and discuss the pros and cons to each. This tutorial leverages the following packages. Learning more: Where you can learn more.Predicting: Apply your final model to a new data set to make predictions.Tuning: Understanding the hyperparameters we can tune and performing grid search with ranger & h2o.Basic implementation: Implementing regression trees in R.The idea: A quick overview of how random forests work.Replication Requirements: What you’ll need to reproduce the analysis in this tutorial.This tutorial will cover the following material: This tutorial serves as an introduction to the random forests. This tutorial will cover the fundamentals of random forests. Random forests are a modification of bagging that builds a large collection of de-correlated trees and have become a very popular “out-of-the-box” learning algorithm that enjoys good predictive performance. Unfortunately, bagging regression trees typically suffers from tree correlation, which reduces the overall performance of the model. Bagging ( bootstrap aggregating) regression trees is a technique that can turn a single tree model with high variance and poor predictive power into a fairly accurate prediction function.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
